Advantages of live-cell imaging
Live-cell imaging in comparison to fixed, endpoint, fluorescent staining-based assays investigates cellular processes over time. This gives more complete information on the dynamics of certain biological processes, as opposed to a single snapshot at a given, and often artificially chosen, time point. Continuous monitoring of cells offers unique advantages:
- It allows the determination of appropriate time points for endpoint studies: many biological processes take a long time (hours to weeks) and often choosing the most significant time points for analysis can be challenging, e.g. when to collect cells after treatment or differentiation, etc. Being able to monitor the cells’ response in real-time enables more accurate identification of relevant endpoints
- It provides additional and continuous data points of biological processes over time, giving more robust quantitative analysis
- It enables quantification of transient phenotypic changes in cells that could be missed if only looking at given time points
- It helps to improve the interpretation of conflicting results from endpoint studies that can be an outcome of reversible processes that are dynamic and controlled in space and time
The emergence of fluorescence tools has started a new era in biological research and enabled us to see and study cellular processes and dynamics that were not possible before. Fluorescent dyes and protein tags, such as green fluorescent protein (GFP), allow monitoring of certain biological processes and dynamics at single-cell resolution and thus enable us to obtain data on heterogeneity that would not be possible with other methods relying on harvesting a population of cells.
--------------------------------------------------------
With CytoSMART Lux3 FL fluorescence cell imager, researchers can track dynamic cellular processes with high specificity by taking high-quality images and real-time time-lapse movies. Learn more about the Lux3 FL here.
--------------------------------------------------------
Key applications of live-cell fluorescence microscopy
The combination of fluorescence and live-cell imaging has become a crucial instrument for modern cellular biology. There are countless possibilities to harness the combined power of both methods. Below we are going to discuss several key applications (Figure 1).
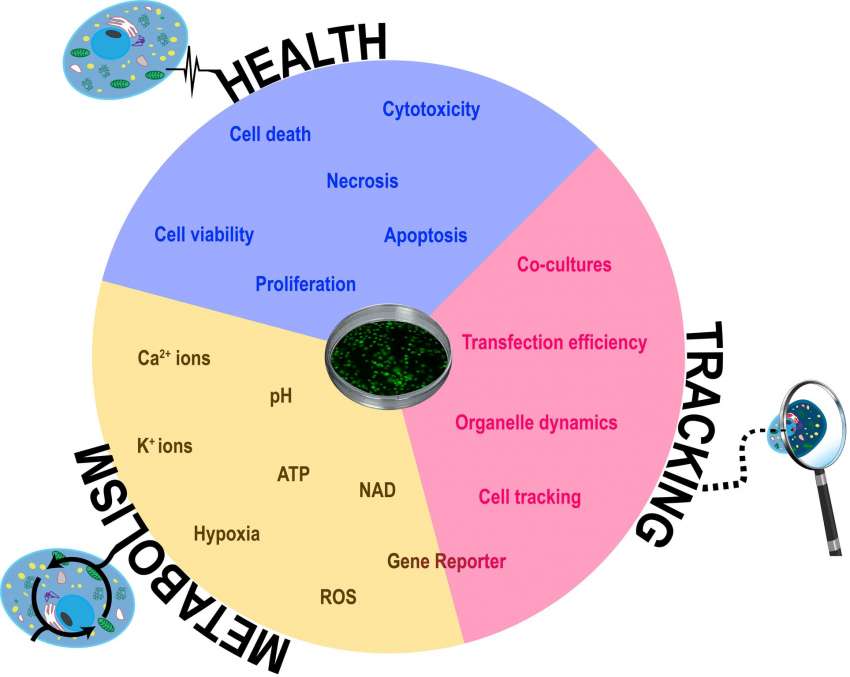
Cell health and viability
Measuring cell health and viability is one of the basic tasks in many areas of biological research including analyzing the effect of different drug treatments, environmental changes, or any other perturbations to the physiological system. Different fluorescent probes allow us to measure different parameters of cell health in real-time. These include cytotoxicity, apoptosis, and proliferation assays.
The basic principle of the majority of assays measuring viability is assessing plasma membrane integrity as the main parameter to identify dead cells. For this purpose, DNA binding fluorophores, which cannot cross cell membranes of living cells and will fluoresce only when bound to nucleic acid, are used. Cells that are not healthy or are dying have compromised membranes that allow for the influx of the dye into the cell where it will bind to the exposed DNA generating fluorescence that can be detected. This serves as an indicator of the viability of individual cells in the population. The most commonly used dyes include Propidium Iodide (PI) or Ethidium Homodimer I dye (EthD-I). These can be multiplexed with other dyes such as Hoechst 33324 to detect the total number of cells or Calcein AM to identify live cells.
Calcein AM is an example of a fluorescent dye that measures viability by identifying all the live cells in culture. In itself, calcein AM is a non-fluorescent, hydrophobic compound that can easily permeate the plasma membrane. In the cytoplasm, it gets cleaved by endogenous esterases producing a strongly fluorescent compound calcein that is retained in cells with intact membranes. Since the fluorescence signal is proportional to the number of live cells, it can also be used to evaluate the number of live cells over several days and thus also serve as a method to assess cell proliferation rate.
Video 1. Live-dead imaging in spheroid cultures. Live cells are stained green and dying cells in the core of the spheroid appear as red.
Some fluorescent dyes allow us to distinguish between different stages of cell death including early apoptotic cells and necrotic cells. Dyes such as oxazole yellow (also known as Yo-Pro1) are high-affinity carbocyanine monomeric nucleic acid stains. Early apoptotic cells become permeable to this compound, while they still remain impermeant to PI (necrotic cells).
Studying apoptosis pathway with fluorescence imaging
Ability to distinguish how certain treatments or compounds can activate the different cell death pathways is essential not only for drug discovery research but also for a better understanding of various biological processes in cells. There are constantly new fluorescent stains being developed to help researchers analyze the different pathways associated with cell death.
One method utilizes the caspase3/7 enzyme system to visualize the apoptotic cells. DNA fluorogenic dyes that have been coupled to the caspase-3/7 recognition sequence can permeate the cell membrane. During apoptosis when the caspase 3/7 is activated, it cleaves off the recognition sequence and thus releases the fluorophore, which is then free to bind DNA leading to fluorescent staining. Another method detects the translocation of phosphatidylserine (PS) from the inner to the outer layer of the plasma membrane as a marker of apoptosis. Annexin V is a PS binding protein that can be conjugated to new-generation fluorophores that are brighter and more stable, allowing visualization of this process in cell cultures (for example see Video 2).
Video 2. Live imaging of cell death induced in CHO cells. Early apoptotic cells turn green (pSIVA) and later when the cell membranes disintegrate (necrotic cells) they turn red (PI).
Measuring cell proliferation with fluorescence imaging
More complex genetically encoded fluorescent probes (GEFPs) enable us to also monitor cell proliferation more in detail and dissect different stages of the cell cycle and division using the fluorescent ubiquitination cell-cycle indicator (FUCCI) developed by Miyawaki et al [1]. This clever system takes advantage of two oscillating proteins hallmarking specific cell cycle transitions. The two-colored probes label G1 phase nuclei red and those in S/G2/M phases green, with the cells in the transition phase from G1 to S appearing yellow. This method thus permits a new level of spatial and temporal resolution to study cell cycle event coordination in response to various treatments or physiological events.
Co-cultures and cell tracking
More complex culture models including 3D cultures such as spheroids and organoids or multi-cell type co-cultures are becoming increasingly important and widespread as in vitro models. Fluorescent labels permit us to track the fate of certain cell subpopulations in a more complex culture and determine how they self-organize, interact with other cell types, etc. Cells can be transfected to express genetically encoded fluorescent labels that are being continuously synthesized and are therefore more suitable for longer-lasting experiments. They require an extra step of transfection and cell selection to get the appropriate line. For short-term experiments or studies where transfection is not a viable option, there are also other available “trackers”.
Some cell-permeable fluorescent dyes such as CMFDA and its variants are nontoxic and retained by cells, staining the whole cytoplasm of cells. Another option is dyes like PKH26 or PKH67 that stain the cell membrane by incorporating aliphatic reporter molecules into the cell membrane lipid bilayer. However, most of these will dilute out with each cell division, therefore the number of days that imaging is possible depends on the speed of cell proliferation.
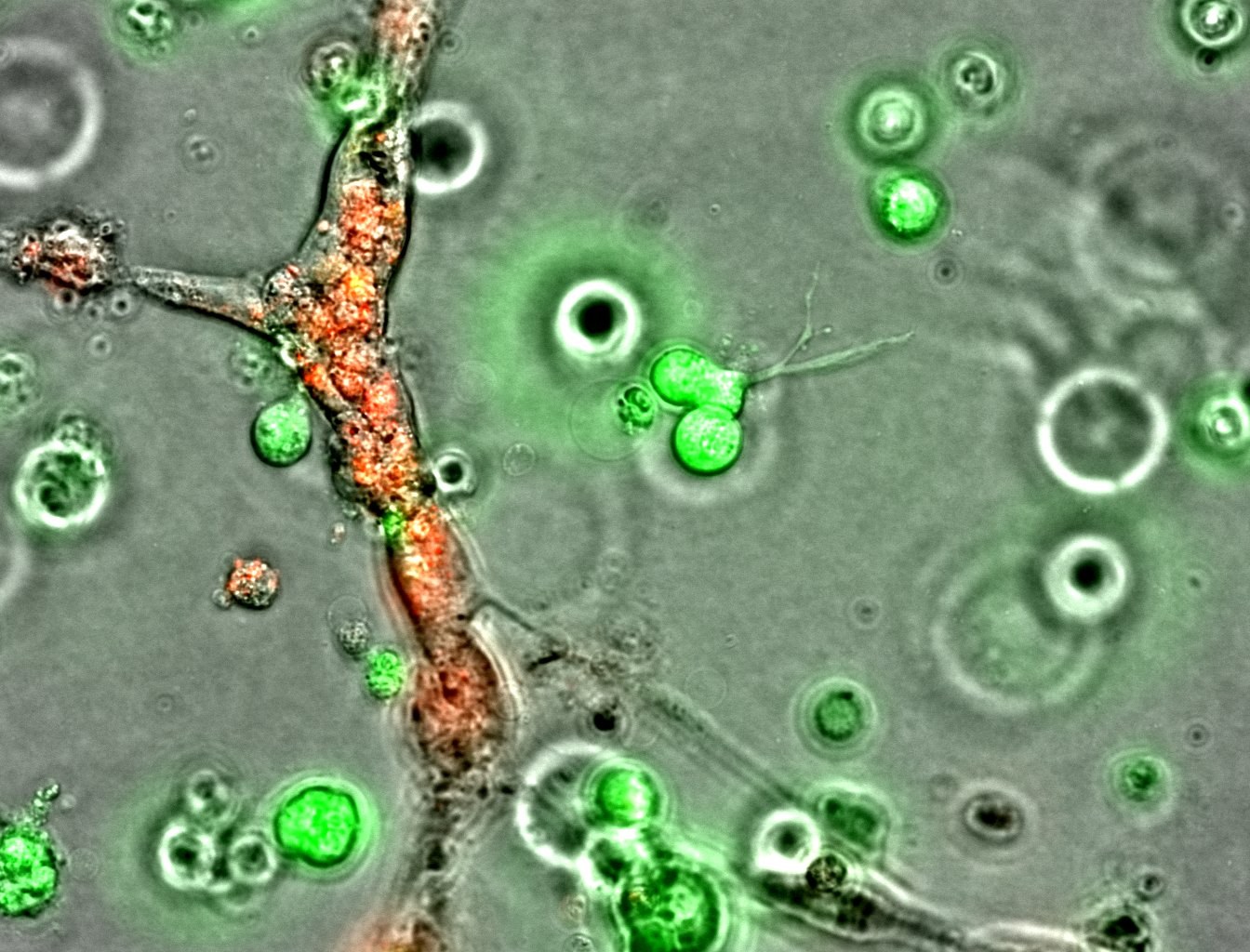
Individual cell tracking with fluorescence imaging
Fluorescent labels enable us to track individual cells - not only their location but also migration and associated changes in morphology. Fluorescent labeling also allows us to see and track cells that are grown on non-transparent materials, such as different scaffolds or hydrogels, for example, in tissue engineering or biomaterials research applications.
Organelle-specific fluorescent probes enable us to track processes within individual cells by visualizing different organelles and their dynamics over time. The probe portfolio now includes labels not only for basic structures, such as the nucleus, cytoplasm, and cell membrane, but also specific ones for labeling mitochondria, cytoskeletal proteins (tubulins and actin), and lysosomes harnessing the organelle-specific properties (i.e. membrane potential, structure, pH) to selectively stain them. They can be used to study cellular compartments in real-time in response to biological events or in combination with other tags to confirm the location or identity of specific proteins.
Studying transfection with fluorescent tags
Transfection has become one of the basic methods of modern molecular biology. During transfection foreign genetic material (DNA or RNA) is introduced into cells to study the function and regulation of genes and proteins.
One of the key checkpoints of any transfection protocol is evaluating the efficiency of this process. Many plasmid constructs, therefore, contain fluorescent protein (GFP, RFP, mCherry, etc) tags or reporters for visualization of the efficacy of this process. With live imaging, it becomes very easy to follow the time-course of expression of the protein of interest as well as compare the efficiency of different transfection conditions to optimize the experimental setup.
Video 3. Time-course of green fluorescent protein (GFP) expression in 3T3 cells transfected with CellLight Nucleus-GFP.
Real-time kinetic data of the experiments
Fluorescent reporters in combination with live imaging offer a plethora of possibilities for experimental design. They can be used to observe and record the activation of specific signaling pathways and produce real-time kinetic data, as well as to quantify the expression of a gene of interest. Cloning the fluorescent reporter downstream of regulatory elements such as promoters can be used to measure their activity.
In comparison to luciferase assays, a traditional endpoint type of experiment, live imaging can produce real-time kinetic data. If these regulatory elements coupled to a fluorescent reporter gene are stably transfected into cells, they can generate specific promoter-reporter cell lines. These are very useful, for example, in stem cell research. Fluorescence reporters that are expressed after activation of certain promoters during stem cell differentiation can provide essential data on the efficiency of differentiation protocols, the time course of different gene expression during the experiment, or within different subpopulations of cells present in the culture [2].
Cell metabolism and fluorescence live-cell imaging
The ongoing development of new probes is continuously expanding the repertoire of parameters that we can monitor in live cells. The last couple of years has seen a significant expansion in the commercialization of new dyes to follow signaling and metabolic processes. These experiments allow us to get a better spatial and temporal resolution of cellular events.
For example, hypoxia dyes are used to monitor oxygen levels in cells and increase fluorescence intensity with decreasing oxygen levels. This can be a very powerful tool in monitoring the health and hypoxia development in 3D cultures, such as spheroids and organoids, where low oxygen concentration levels can lead to the formation of a necrotic core, or in studying the effects of hypoxia on cell metabolism. Other probes are able to visualize oxidative stress in live cells, as they emit fluorescence when they are oxidized by reactive oxygen species (ROS). Oxidative stress is implicated in many biological processes, such as senescence, inflammation, and certain degenerative diseases, and arises due to the generation of ROS in cultures and when cells are unable to sufficiently scavenge it.
The scope of possible applications has been expanded further by a wide range of GEFPs. They are usually constructed by a fluorescent protein and a selective domain that either recognizes a certain analyte or is modified by a signaling process that in turn changes the fluorescent properties of the reporter. Some examples include intracellular calcium probes (GECIs), potassium ion probes (GEPPIs), intracellular pH indicators, ATP-sensors, and many more. Furthermore, the probes can be now designed to be non-specific or to target certain organelles (e.g. mitochondria, ER, lysosomes, etc.). For an extensive review see Depaoli et al (2019) [3].
How to set up a fluorescence live-cell imaging experiment
Live-cell fluorescence imaging has many advantages and offers a variety of experimental design opportunities. However, as with any other method, there are some considerations that need to be taken into account when setting up the experiment to obtain meaningful data and to avoid damage to the cells and potential artifacts.
Keep cells in perfect conditions
First of all, cells need to be maintained in a physiological environment throughout the whole duration of the experiment. Some immortalized cell lines are robust enough to be imaged for the short term without any environmental control. However, changes in temperature, oxygen, and carbon dioxide (CO2) concentration can lead to changes in cell behavior and impact the observed processes. Therefore, in longer experiments, the imaging system with a humidified heated chamber and CO2 control can be used. However, fluorescence live-cell imaging devices that fit and work inside an incubator are preferred.
Choose the right fluorescent dye
The choice of the dye is equally important. Genetically encoded fluorescent proteins are more specific and stable over time in culture; however, they do imply genetic manipulation of the cells including all the possible negative effects that this process can have on cell physiology. In the case of the chemical fluorophores, their effects on cells in terms of cytotoxicity, permeability, and retention over time need to be considered. The selection will be obviously also influenced by the excitation and emission spectra of the probe depending on the fluorescence microscope filter setup.
Minimize the light intensity
Lastly, there are negative effects in cells caused by their exposure to high-intensity excitation light during the imaging process. The photon flux is damaging to cells by creating excessive amounts of reactive species, such as free radicals and ROS, in a process called phototoxicity. Therefore it is desirable to design experiments in such a way as to minimize the light (wavelength range and number of photons) that cells are exposed to, but at the same time to optimize the signal to noise ratio. This depends not only on the brightness of the fluorescent label itself but also on the light source of the fluorescent microscope (LED lights vs mercury lamps).
Mind non-invasiveness
All in all, awareness of the notion that fluorescence live microscopy is not entirely non-invasive can help us design experiments to minimize cell harm, non-physiological responses, and experimental artifacts and at the same time obtain detailed information about cellular processes over time.
Conclusion
The latest advances in the field of live-cell fluorescence microscopy and the ever-growing range of both genetically encoded fluorophores and chemical dyes have created many new opportunities in the fields of cellular and molecular biology. Moreover, the possibility to multiplex the different assays creates a scope for many innovative techniques and experiments that will provide a better understanding of cell behavior and the dynamics of underlying molecular processes in real time.
Related Products
There are currently no products tagged to this resource.