Cell migration is involved in a multitude of critical physiological and pathological processes. Cell motility can be divided into collective and single-cell migration1. Single-cell migration is used by cells to move towards and between tissues, and it can be split into amoeboid and mesenchymal migration2,3. A variety of in vitro assays have been developed to study single-cell migration.
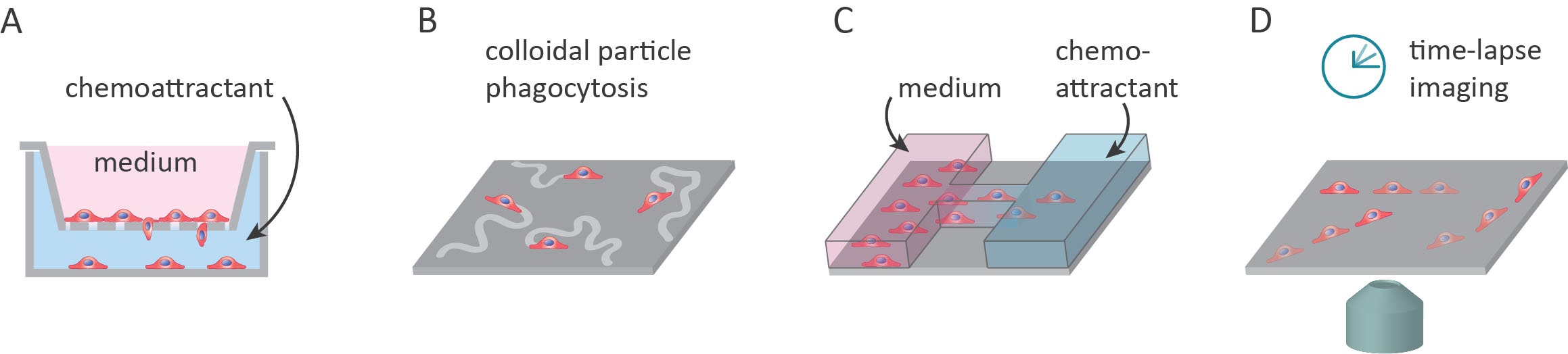
Several factors are essential in the design of single-cell migration assays. These factors include robust quantification, the use of a few step protocols, and a close resemblance to the cell microenvironment found in vivo. In addition to these factors, assays that use three-dimensional (3D) culture environments and can accommodate the co-culturing of multiple cell types can also be desirable4. This article will cover the most well-known methods for investigating two-dimensional (2D) single-cell migration, these are summarized in Figure 1.
1. Boyden chamber assay
The Boyden chamber assay also referred to as the Transwell assay, is a standard method used for migration analysis5. The Boyden chamber assay is an endpoint assay and indicates the number of cells that have migrated through a membrane.
The setup consists of two stacked culture compartments separated by a porous membrane. Cells are seeded in the upper compartment, and culture medium supplemented with a chemo-attractant is added to the lower compartment. The chamber is then incubated, allowing cells to adhere and form a cell monolayer. Cells can then migrate through to the lower surface of the membrane. After incubation, the cells are fixed and stained. Cells that remain on the upper surface of the membrane are removed using a cotton swab. The stained migrated cells can then be counted using a microscope, or the stain can be resolubilized and quantified5,6. Chambers that exclude light can be used with fluorescent dyes, and this removes the need to count cells and reduces the amount of steps in the protocol7.
The benefit of this assay is that it allows the researcher to study 3D cell chemotaxis and it can be used for both migration (uncoated membrane) and invasion assays (coated membrane)5. A design limitation of the method is that the endpoint result is based upon a complete sample response; therefore, heterogenous populations cannot be distinguished4. Furthermore, migrating cells cannot be imaged in real-time8. An additional drawback is that the assay can be time-consuming as optimization is required to achieve significant differences between experimental groups4. Commercial Boyden chamber assays are available, including Boyden Chamber Assays from Cell Biolabs, Inc., and Transwell and FluoroBlok systems from Corning.
2. Colloidal particle assay
Another assay used to study single-cell migration is the colloidal particle assay, also known as the colloidal gold phagokinetic track assay—this is a straightforward method for observing cell migration. The protocol involves monitoring tracks made by motile cells on specially coated surfaces.
Microplates are first coated using colloidal gold or quantum dots. After that, cells are added at a low cell density to the surface9–11. Using cells at low density prevents trail overlaps10. Motile cells will then phagocytize the coating as they move, creating a phagokinetic track free of colloidal gold or quantum dots. This trail can be visualized under a light microscope, and the area of the path can be compared between different experimental groups10–13.
The advantages of this assay are that cells that are unable to pass through the Boyden chamber assay membranes can be tested, cells can be imaged in real-time, the assay has higher sensitivity compared to the Boyden chamber assay, and the assay is automatable10,13. The limitations of the method are that the preparation of the coating requires optimization for best results, cells that backtrack on the path are not recorded, and is not suitable for the measurement of bulk chemotactic effect, 3D migration, or the detailed investigation of a cellular movement11. No commercial kits are available for this assay.
3. Microfluidic chips
A more sophisticated method for investigating cell migration makes use of custom microdevices and microfluidics8. Microfluidic assays are flexible and can be used for either endpoint or continuous analysis4. These assays allow the researcher greater control of the experimental design and migration channel8. These microdevices usually consist of a silicon substrate fabricated of polydimethylsiloxane (PDMS)4,8. These devices can also include gelatin or other extracellular matrix-based polymers to reflect the cell microenvironment better4. For visualization purposes, cells can also be stained with markers such as green fluorescent protein (GFP)8.
The benefits of microfluidic assays are that they require minimal volumes and provide the researcher with precise microenvironment control8,14. Low volumes are particularly useful when examining small tumor biopsy samples. These devices compare well with Boyden chamber assays in terms of resolution, precision, and investigating chemotaxis4. A disadvantage of microfluidic devices is that they generally do not allow for distinguishing between individual cells, nor do they allow for the retrieval of cells after the migration assay8. An additional drawback of microfluidic chips is that they can be time- and cost-consuming because of microfluidic pumping systems and continuous recording under cell culture conditions. The time and cost can hinder its high-throughput capability and the use of this technique by researchers4. Commercial kits include the Millicell® µ-Migration Assay Kit from Merck and the µ-Slide Chemotaxis by ibidi.
4. Time-lapse cell tracking
This technique allows researchers to track individual cells in real time15. In this protocol, cells are seeded on a surface and allowed to adhere. This surface is mounted onto an environmentally controlled microscope stage or on an in-incubator microscopy system such as the CytoSMART Lux2. After adhesion, images of the migrating cells are recorded at five to ten-minute intervals6,15,16. These time-lapse movies are used to identify and track single cells that are not undergoing cell division. This technique can be combined with microfluidic devices.
The advantage of live-cell time-lapse video microscopy followed by cell tracking is that other cell migration methods often only report on endpoint results and can be influenced by cell proliferation and cell-cell contacts6. Furthermore, not only the migratory path can be determined using this technique, but also the migration speed and persistence. Commercial software for single-cell monitoring exists such as MetaMorph by BioImaging Solutions Inc, and there are many free to download software that can be used, such as TrackMate by ImageJ. A comparative list of available software available to analyze single-cell migration data is summarized by Massuzo et al. (2017)17.
--------------------------
For researchers looking to perform cell tracking assays using time-lapse imaging, we recommend the CytoSMART Lux2 Duo Kit. This mini live-cell imaging microscope fits inside cell culture incubators and can monitor cells in culture vessels and microfluidic chips.
--------------------------
5. Conclusion
In this section, commonly used assays investigating single-cell migration were discussed. The methods covered were the Boyden chamber assay, colloidal particle assay, microfluidic chips, and time-lapse cell tracking. These techniques include both endpoint and continuous analysis and can be used to quantify both the whole sample and individual cell responses. In addition to single-cell migration, cells can migrate collectively. Information relating to collective cell migration techniques, specifically cell removal, exclusion, and outgrowth assays, form part of this series of articles on cell migration.
| Method | Advantages | Disadvantages | Commercial products |
References |
|---|---|---|---|---|
| Boyden chamber assay | 3D cell chemotaxis, migration/invasion assays | Endpoint analysis, no real-time imaging, time-consuming, not all cell type compatible | Boyden Chamber Assays (Cell Biolabs, Inc.), Transwell, and FluoroBlok systems (Corning) | 5 |
| Colloidal particle assay | Real-time imaging, automatable | The coating requires optimization, no 3D migration, cell backtracking not measured | - | 11 |
| Microfluidic chips | Minimal volumes, precise microenvironment control, endpoint/continuous analysis, customizable | Time-consuming, expensive, individual cells not usually analyzed | Millicell® µ-Migration Assay Kit (Merck), µ-Slide Chemotaxis (ibidi) | 4, 8 |
| Time-lapse cell tracking | Continuous analysis, no cell proliferation, no cell-cell contacts | Data-intensive | MetaMorph (BioImaging Solutions Inc), TrackMate (ImageJ) | 15, 17 |
Related Products
There are currently no products tagged to this resource.